AutoSciDACT: A Universal Pipeline for Automated Scientific Discovery
- The Challenge: Finding Needles in Scientific Haystacks
- Why Existing Anomaly Detection Falls Short for Science
- The AutoSciDACT Pipeline: Two Phases
- Experiments Across Four Scientific Domains
- Key Results: Sensitivity Across Domains
- Bonus: Re-Discovering the Higgs Boson in Real LHC Data
- Why This Matters: Bridging AI and the Scientific Method
- References
Samuel Bright-Thonney, Christina Reissel, Gaia Grosso, Nathaniel Woodward, Katya Govorkova, Andrzej Novak, Sang Eon Park, Eric Moreno, Philip Harris
MIT Department of Physics & The NSF AI Institute for Artificial Intelligence and Fundamental Interactions (IAIFI)
Published at NeurIPS 2025 | OpenReview
The Challenge: Finding Needles in Scientific Haystacks
Scientific discovery has always had an element of serendipity. The cosmic microwave background was first noticed as unexplained antenna noise. Penicillin emerged from a contaminated petri dish. X-rays were found while studying cathode rays. In each case, a curious anomaly in data – something that didn’t fit the expected pattern – led to a breakthrough.
Today’s scientists face a version of this challenge at an unprecedented scale. Modern experiments generate torrents of data: the Large Hadron Collider (LHC) produces roughly a petabyte per second of raw collision data; gravitational wave observatories continuously stream time-series from multiple detectors; biomedical imaging pipelines churn out terabytes of high-resolution tissue scans. Somewhere in these massive, noisy, high-dimensional datasets, there may be genuine novelties waiting to be found – a new particle, an unexpected astrophysical signal, an early marker of disease. But how do you find something you don’t know you’re looking for, and how do you prove it’s real?
This is the problem AutoSciDACT (Automated Scientific Discovery with Anomalous Contrastive Testing) was built to solve. In our NeurIPS 2025 paper, we introduce what is – to our knowledge – the first end-to-end, domain-agnostic pipeline for novelty detection in scientific data that provides rigorous statistical guarantees. AutoSciDACT doesn’t just flag anomalies; it tells you how confident you should be that what you’ve found is real.
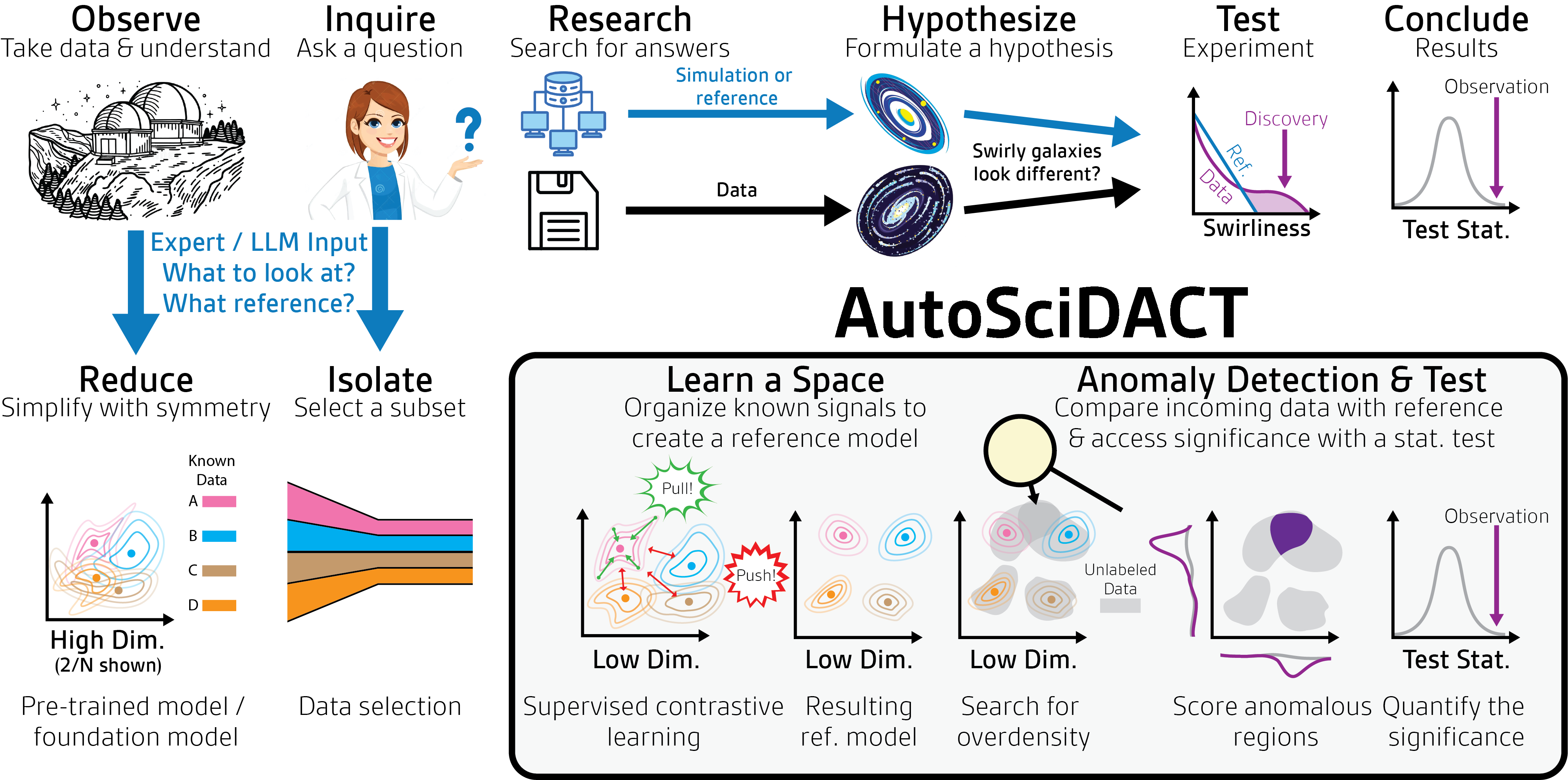
Why Existing Anomaly Detection Falls Short for Science
The machine learning community has developed a rich toolkit for anomaly and out-of-distribution (OOD) detection. Methods like autoencoders, contrastive OOD detectors, and one-class classifiers are effective at flagging individual data points that look “weird.” But science demands more than a list of suspicious data points. It demands a p-value – a quantification of how likely the observed data is under the assumption that nothing new is going on (the null hypothesis).
Consider the difference:
- Standard anomaly detection says: “These 50 data points look unusual.”
- Scientific hypothesis testing says: “The probability of seeing this pattern of anomalies if nothing new is happening is less than 1 in 10,000.”
This distinction matters enormously. In particle physics, claiming a “discovery” requires evidence at the 5-sigma level (a p-value of about 3 in 10 million). In other fields, the bar may be lower, but the principle is the same: you need a statistical framework that accounts for the look-elsewhere effect, the trial factors, and the noise characteristics of your experiment. Most ML-based anomaly detectors simply aren’t designed for this.
There’s a second, equally fundamental challenge: dimensionality. Raw scientific data is often extremely high-dimensional – a single jet measured by an experiment at the LHC might consist of hundreds of particles, each with 10 or more measured features; a gravitational wave observation is a multi-channel time series with thousands of samples; a histology image has thousands of pixels. Statistical tests lose power rapidly as dimensionality grows (the curse of dimensionality), so any practical pipeline needs to compress data into a compact, meaningful representation before testing.
The AutoSciDACT Pipeline: Two Phases
AutoSciDACT addresses both challenges with a clean two-phase design: pre-training (learn a compact representation) followed by discovery (test for anomalies with statistical rigor).
Phase 1: Contrastive Pre-Training
The first phase uses supervised contrastive learning to compress high-dimensional scientific data into an expressive 4-dimensional embedding. The key idea is simple but powerful: use labeled examples of known phenomena to teach an encoder what “normal” looks like, so that anything truly new will stand out in the learned space.
The backbone of the pipeline is an encoder \(f_\theta : \mathcal{X} \to \mathbb{R}^d\) that maps raw data from its high-dimensional input space \(\mathcal{X}\) to a compact \(d\)-dimensional representation. We train this encoder using the SupCon (supervised contrastive) objective [1], which pulls together embeddings of same-class data and pushes apart embeddings of different classes. This is a generalization of the self-supervised SimCLR framework [2] that uses class labels rather than data augmentations to define positive pairs. Given a batch \(\mathcal{B}\) of inputs, the loss is:
\[\mathcal{L}_{\text{SupCon}} = -\sum_{i \in \mathcal{B}} \frac{1}{|P(i)|} \sum_{p \in P(i)} \log \frac{\exp\bigl(\text{sim}(\mathbf{z}_i, \mathbf{z}_p) / \tau\bigr)}{\sum_{j \neq i} \exp\bigl(\text{sim}(\mathbf{z}_i, \mathbf{z}_j) / \tau\bigr)}\]Here \(\mathbf{z} = g_\phi(f_\theta(\mathbf{x}))\) is the output of a small projection head \(g_\phi\) applied to the encoder embedding, \(\text{sim}(\cdot, \cdot)\) is cosine similarity, \(\tau\) is a temperature parameter, and \(P(i)\) is the set of all samples in the batch sharing the same class label as sample \(i\). The numerator rewards pulling same-class embeddings together; the denominator penalizes failing to separate different classes. After training, the projection head is discarded and we use \(\mathbf{h} = f_\theta(\mathbf{x})\) as the embedding for downstream tasks.
In practice, we augment this with a sub-dominant cross-entropy classification loss, giving a total objective \(\mathcal{L} = \mathcal{L}_{\text{SupCon}} + \lambda_{\text{CE}} \mathcal{L}_{\text{CE}}\) (with \(\lambda_{\text{CE}} \sim 0.1\)–\(0.5\)), which we found encourages more regular cluster structure in the embedding space.
Crucially, this approach leverages a resource that many scientific domains have in abundance: high-quality simulations and expert labels. Particle physicists have extraordinarily detailed Monte Carlo simulations. Astronomers can inject synthetic gravitational wave signals into real detector noise. Histopathologists can label tissue types with high accuracy. AutoSciDACT channels all of this domain knowledge into the embedding, but keeps it strictly separate from the downstream analysis – the test itself knows nothing about what specific anomaly might be present.
A subtle but important point: we fix the embedding dimension to just \(d = 4\) for all experiments. This is an intentionally aggressive compression (from inputs that can have hundreds or thousands of dimensions), chosen to keep the subsequent statistical test tractable. Despite this extreme compression, the contrastive objective preserves enough structure to enable highly sensitive anomaly detection.
Phase 2: Discovery with NPLM
The second phase deploys the New Physics Learning Machine (NPLM) [3, 4], a machine learning-based hypothesis test originally developed for particle physics. NPLM implements a likelihood ratio test – the gold standard of statistical hypothesis testing, rooted in the foundational Neyman-Pearson lemma [5] – but replaces the traditional parametric signal model with a flexible, data-driven alternative.
Here’s the intuition: given a reference dataset \(\mathcal{R}\) of known backgrounds and an observed dataset \(\mathcal{D}\) of unknown composition, NPLM asks: “Is there any way the observed data deviates from the reference, and if so, how statistically significant is that deviation?” The mathematical foundation is the Neyman-Pearson test statistic:
\[t(\mathcal{D}) = 2 \max_{\mathbf{w}} \sum_{x \in \mathcal{D}} \log \frac{\mathcal{L}(x \mid \mathcal{H}_{\mathbf{w}})}{\mathcal{L}(x \mid \mathcal{H}_0)}\]This compares the likelihood of the data under a family of learned alternative hypotheses \(\mathcal{H}_{\mathbf{w}}\) to the null hypothesis \(\mathcal{H}_0\) (that \(\mathcal{D}\) and \(\mathcal{R}\) are drawn from the same distribution). The key insight is that the alternative density is parametrized as a reweighting of the null:
\[p(x \mid \mathcal{H}_{\mathbf{w}}) = p(x \mid \mathcal{H}_0) \, \exp\bigl[f_{\mathbf{w}}(x)\bigr]\]where \(f_{\mathbf{w}}(x) = \sum_{i=1}^{M} w_i \, k_i(x)\) is a mixture of \(M\) Gaussian kernels with learnable coefficients [6]. Rather than specifying what the signal is, the model learns whatever deviation best explains the data. In practice, NPLM is trained as a binary classifier distinguishing \(\mathcal{D}\) (labeled \(y=1\)) from \(\mathcal{R}\) (labeled \(y=0\)), optimizing a regularized weighted cross-entropy:
\[\mathcal{L}_{\text{NPLM}} = \sum_{(x,y)} \left[ w_{\mathcal{R}}(1-y)\log(1 + e^{f_{\mathbf{w}}}) + y\log(1 + e^{-f_{\mathbf{w}}}) \right] + \lambda \sum_{i,j} w_i w_j k_i(x_j)\]where \(w_{\mathcal{R}}\) reweights the reference to match the expected yield under the null hypothesis. Once trained, the test statistic \(t_{\text{NP}}(\mathcal{D})\) is computed from the learned \(f_{\hat{\mathbf{w}}}\), and calibrated by running hundreds of pseudo-experiments (“toys”) – resampling datasets under the null hypothesis to build an empirical distribution of test statistics. The p-value is simply the fraction of null toys with a test statistic exceeding the observed one:
\[p = \frac{1}{|\mathcal{T}_0|} \sum_{t \in \mathcal{T}_0} \mathbb{1}[t > t(\mathcal{D})]\]The p-value can also be estimated asymptotically from the distribution of \(\mathcal{T}_0\), which can be fit to a suitable \(\chi^2\) distribution [6]. This is useful in cases where the deviation between \(\mathcal{D}\) and \(\mathcal{R}\) is large (e.g. \(Z = 5\sigma\)), in which case the number of toys necessary for an accurate empirical estimate would be prohibitively large.
Comparisons with other goodness-of-fit approaches [7] have demonstrated NPLM’s impressive sensitivity to subtle distributional distortions. To further improve robustness, we adopt an extension [8] that runs the test at multiple kernel widths and combines the resulting p-values, mitigating sensitivity to the choice of any single kernel scale.
This is fundamentally different from anomaly scoring methods. NPLM doesn’t try to identify which data points are anomalous – it tests whether the distribution of observed data is consistent with the reference. This makes it sensitive to subtle collective effects: small overdensities, slight shape distortions, or faint excesses that no individual data point would reveal.
Experiments Across Four Scientific Domains
To demonstrate that AutoSciDACT genuinely transfers across fields, we tested it on datasets from particle physics, gravitational wave astronomy, histopathology, and genomics – plus synthetic benchmarks and standard image datasets. In each case, the pipeline follows the same recipe: train a contrastive encoder on known classes, hold out one class as the “anomaly,” then see how small a fraction of anomalous data NPLM can detect.
Particle Physics: Finding the Higgs in Jets
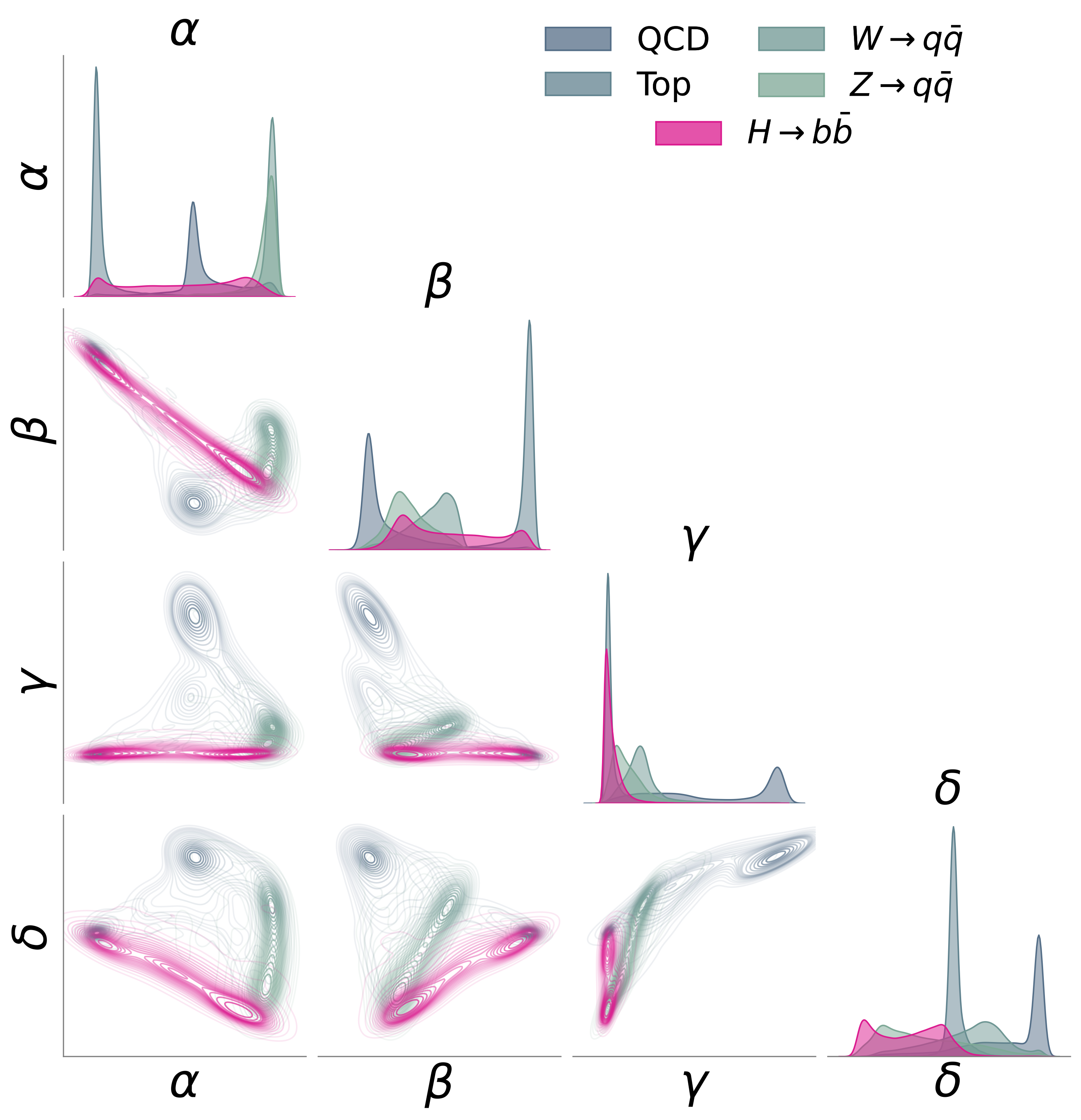
We used the JetClass dataset [9] of simulated particle jets from the LHC – streams of particles produced when quarks and gluons fly apart after high-energy collisions. The encoder (a Particle Transformer [9], a variant of the Transformer architecture [10] adapted for particle physics) was trained on QCD, top quark, and W/Z boson jets. The held-out anomaly: jets from Higgs boson decays to bottom quarks.
AutoSciDACT detected these Higgs jets at over 3-sigma significance with just a ~1% signal contamination in a sample of 10,000 events – approaching the performance of a fully supervised classifier that knows what the Higgs signal looks like.
Gravitational Wave Astronomy: Detecting Unknown Signals in LIGO Data
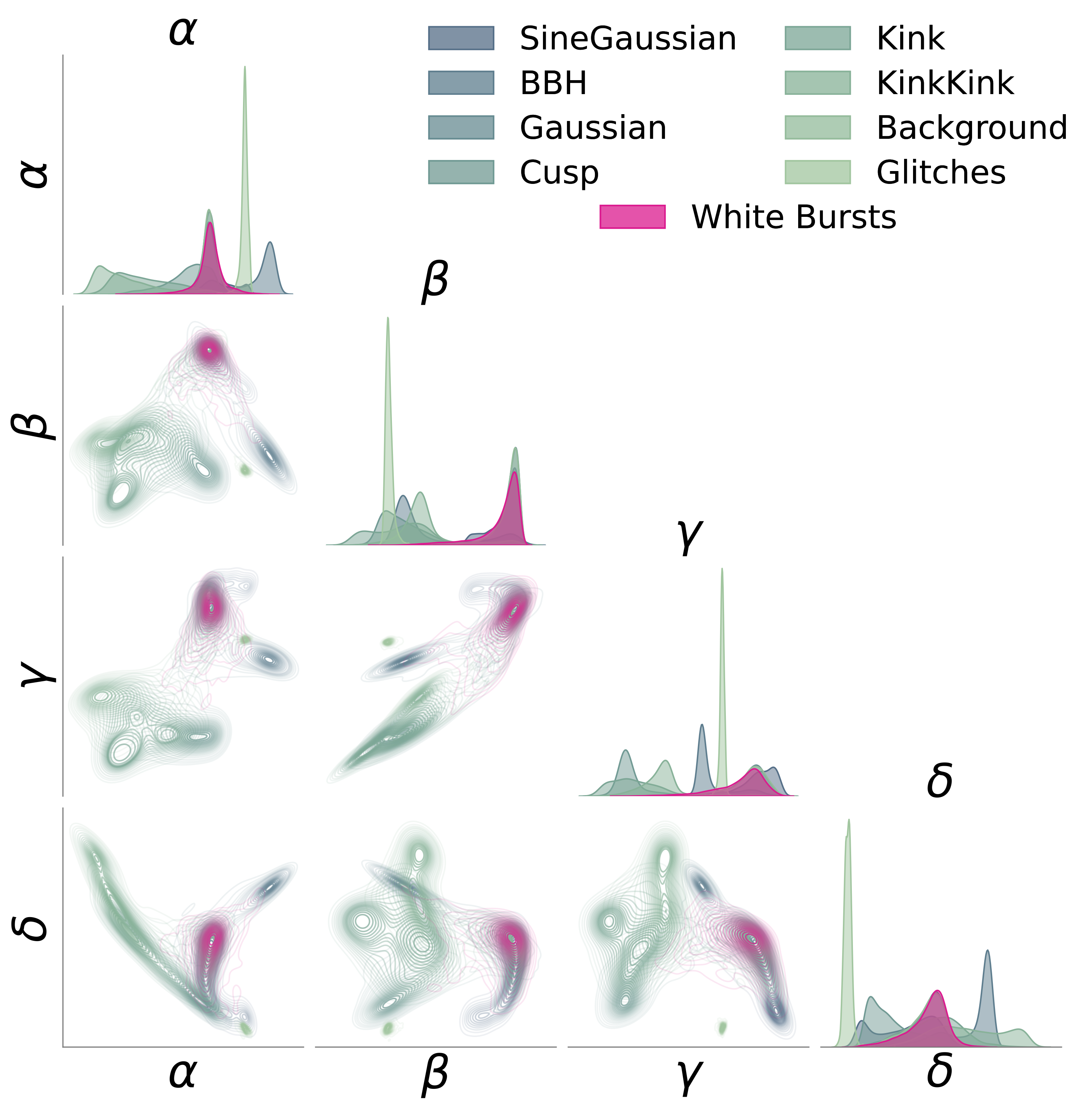
We applied AutoSciDACT to real data from LIGO’s [11] third observing run: 50-millisecond time-series snippets from both the Hanford and Livingston detectors. The encoder (a 1D ResNet [12], following the setup of [13]) was trained on background noise, glitches, and six types of known or hypothetical astrophysical signals. The held-out anomaly: white noise bursts – deliberately model-agnostic signals that lack the distinctive chirp structure of, say, a binary black hole merger. This represents a worst-case scenario for detection: a signal with minimal distinguishing features.
Even so, AutoSciDACT achieved strong statistical significance at low injection fractions, again approaching supervised limits.
Histopathology: Early Disease Detection in Tissue
For the biomedical application, we used histological images [14] of stained tissue samples: seven classes of normal mouse tissue (brain, heart, kidney, liver, lung, pancreas, spleen) plus normal rat liver. The anomaly: mouse liver tissue affected by non-alcoholic fatty liver disease (NAFLD). The encoder was an EfficientNet-B0 [15], following the approach of [16].
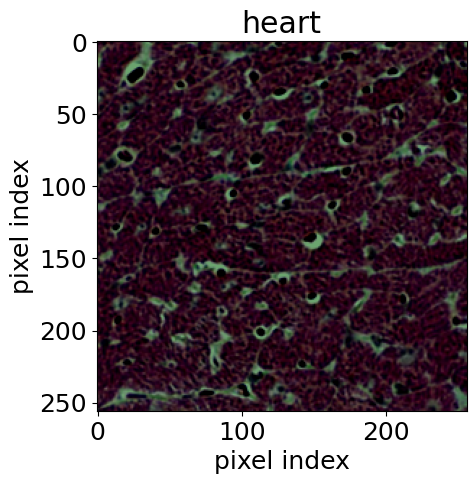
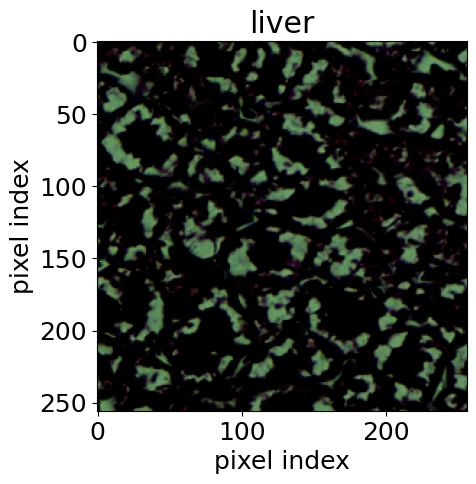
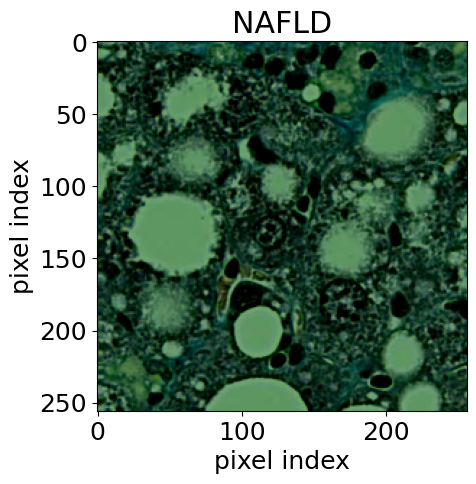
This example is particularly compelling because it demonstrates transfer of methodology across disciplines. Rigorous hypothesis testing of this kind is standard practice in physics, but much less common in pathology. AutoSciDACT brings those techniques to a domain where they could enable earlier, more sensitive detection of disease – especially when only a small fraction of tissue in a sample is affected.
Genomics: Detecting Hybrid Butterflies
We also tested on a genomics-adjacent task: detecting hybrid butterfly species from wing images. Butterfly subspecies have characteristic wing patterns, and hybrids can exhibit subtle blends of their parent species’ features. Using a BioClip-based [17] encoder trained on 14 subspecies, AutoSciDACT successfully detected hybrid individuals even at low injection fractions.



Key Results: Sensitivity Across Domains
The headline result is summarized in the figures below, showing detection significance (Z-score) as a function of the fraction of anomalous signal injected into each dataset. In many cases, NPLM achieves highly significant detections (\(Z \gtrsim 3\)) with small signal fractions.
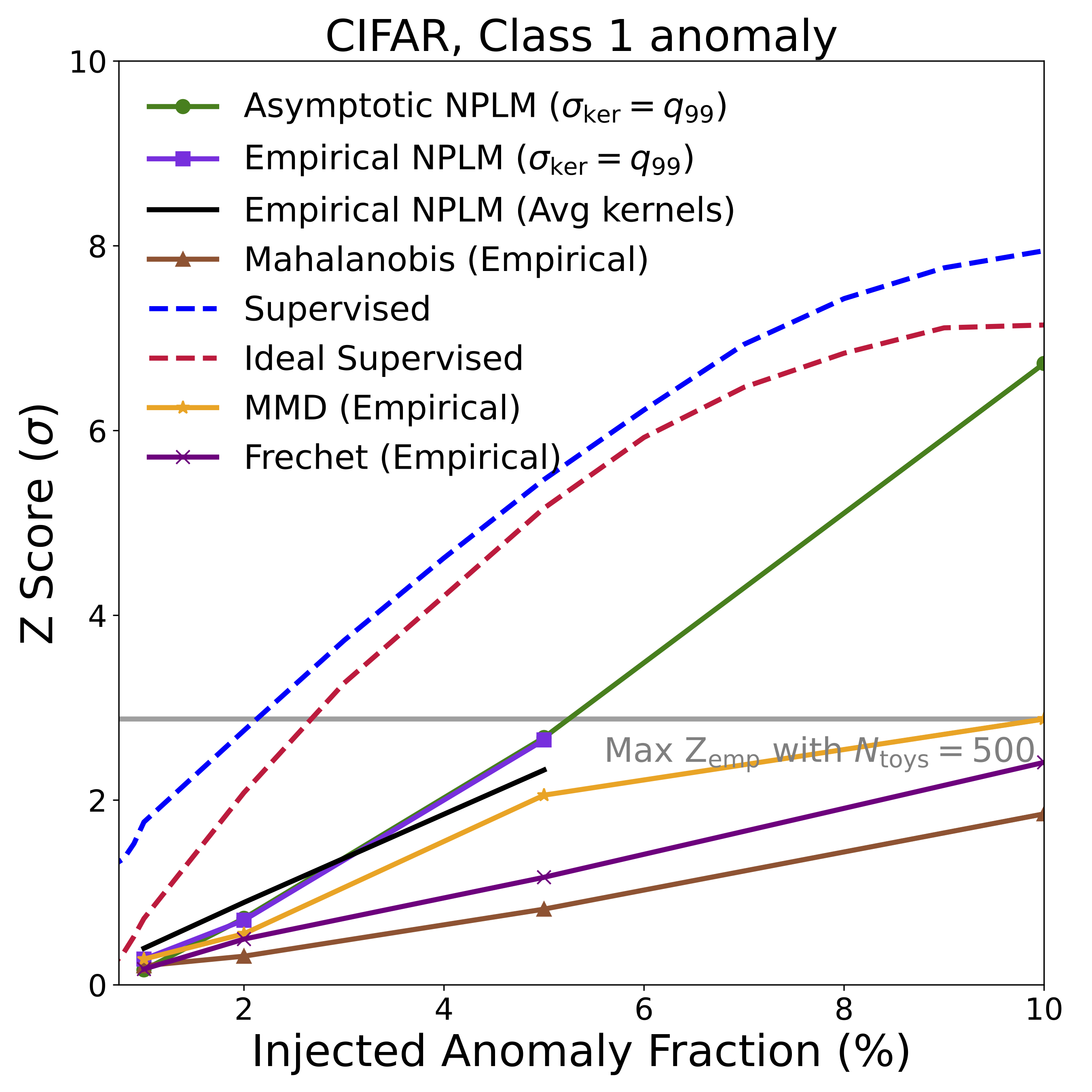
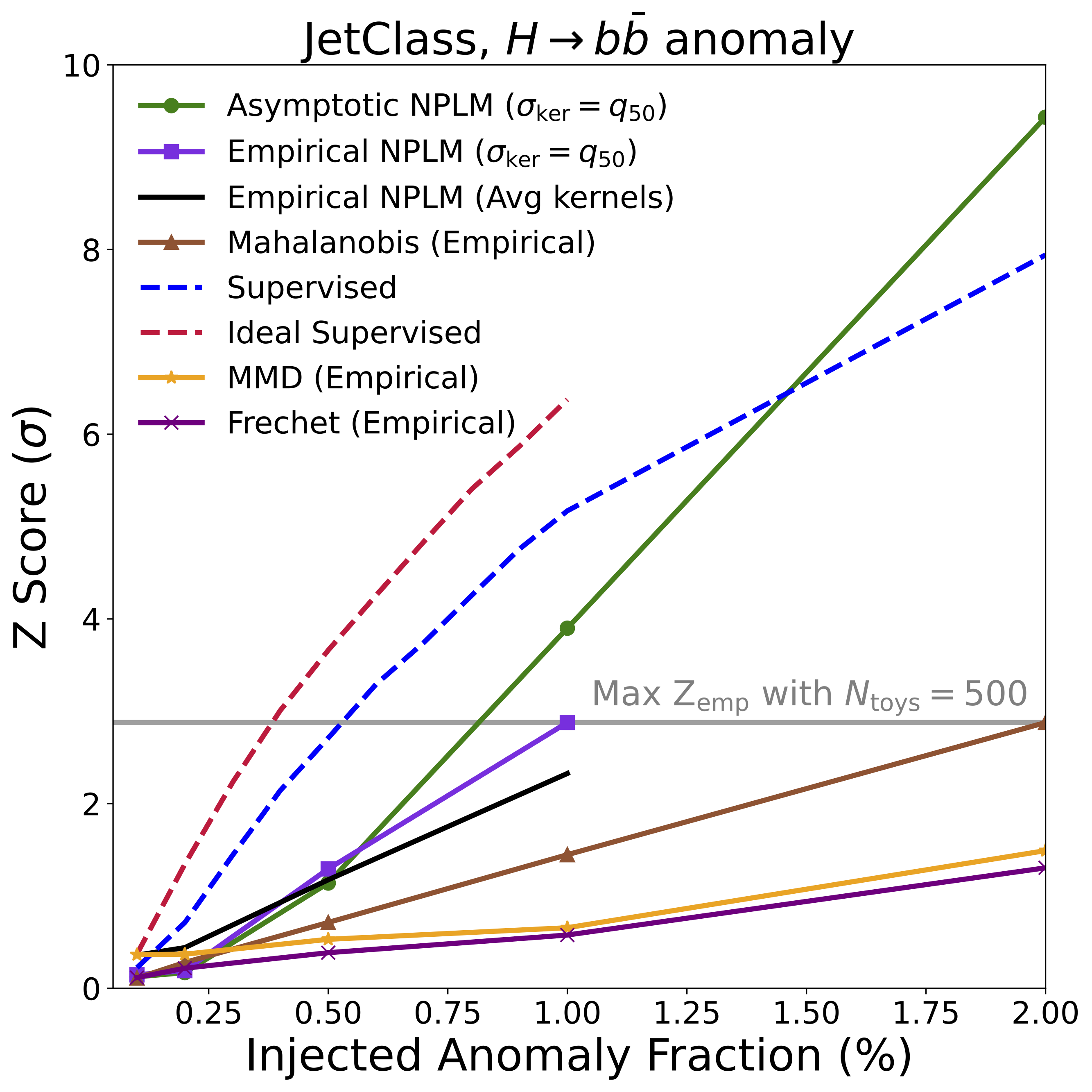
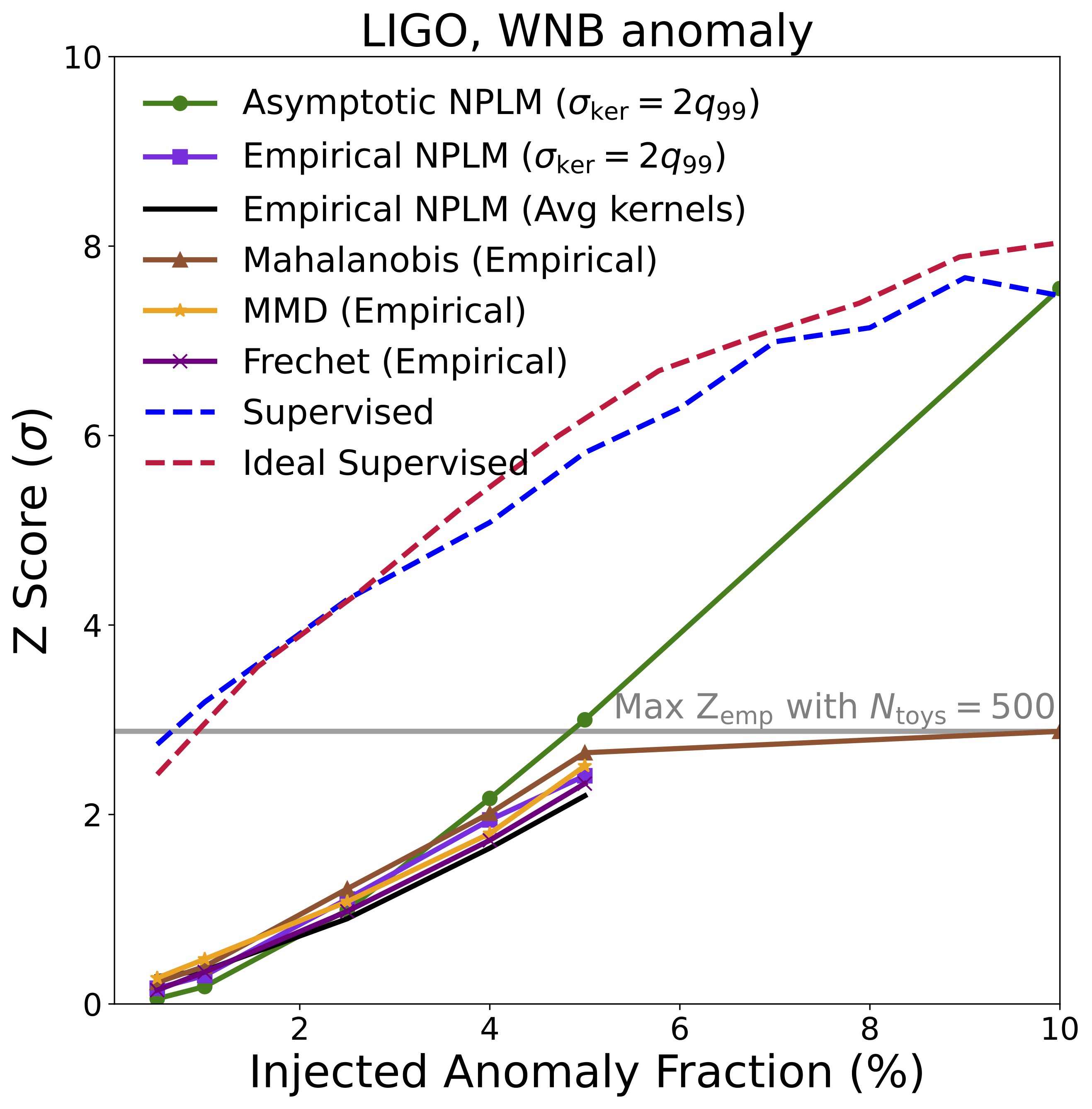
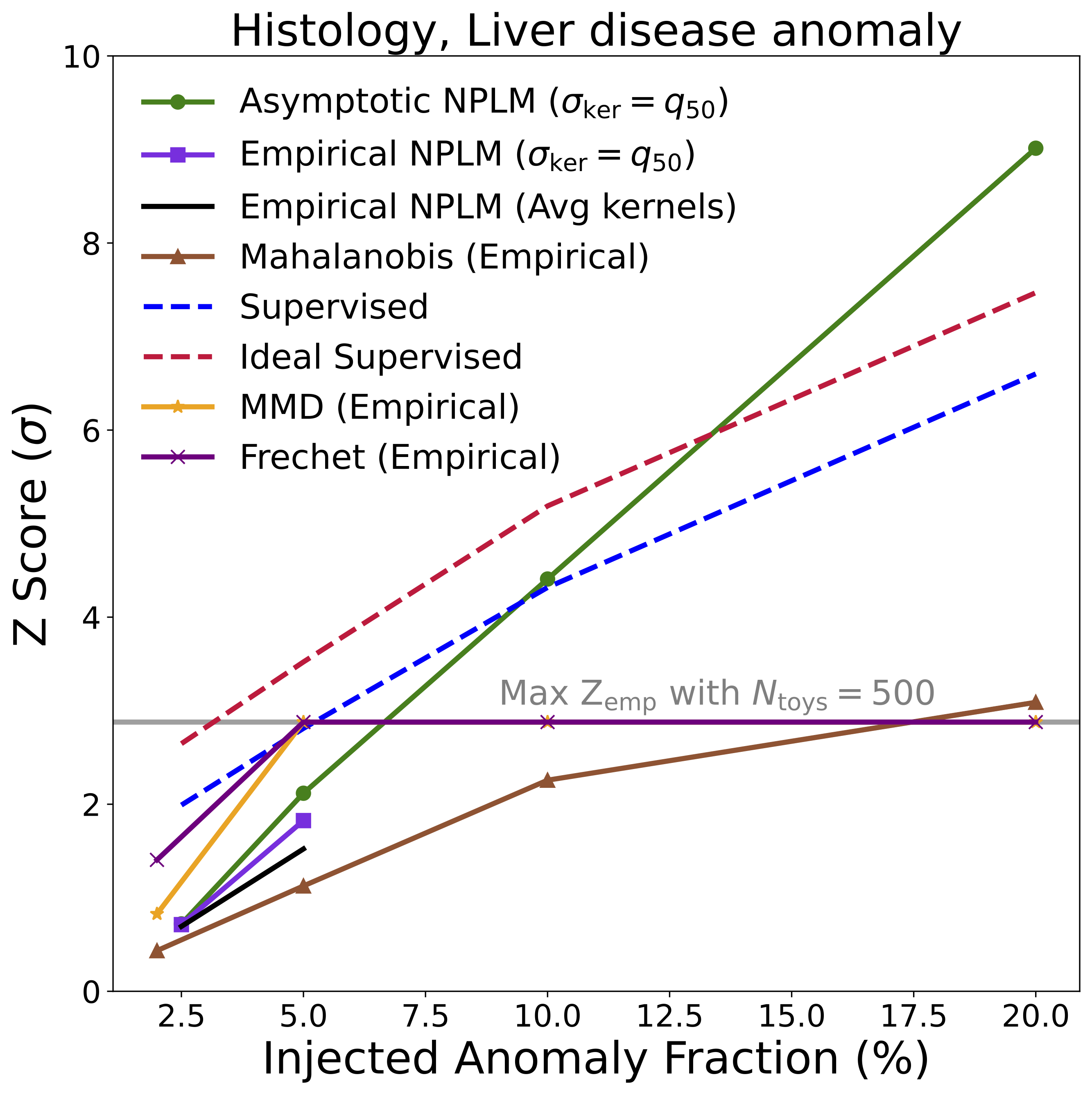
Several things stand out:
-
NPLM performs strongly while remaining signal-agnostic. AutoSciDACT makes no assumptions about the distribution or frequency of anomalies in the observed dataset, meaning it is signal agnostic. In several cases, the unsupervised NPLM test approaches the sensitivity of classifiers that explicitly know what the anomaly looks like. This is remarkable, and speaks to the quality of the contrastive embeddings. For both the LIGO and JetClass datasets, our results rival or exceed existing anomaly detection algorithms within their respective domains [19, 20].
-
The pipeline is robust to high-dimensional noise. In synthetic experiments, we showed that adding dozens of uninformative noise dimensions to the raw data rapidly degrades detection with traditional methods, but has almost no impact on AutoSciDACT thanks to the contrastive compression. (Check out the paper for detailed results!)
-
NPLM outperforms simpler statistical tests. The Mahalanobis distance baseline [18], which assumes Gaussian cluster structure, struggles when anomalies manifest as overdensities near existing clusters rather than isolated outliers. NPLM’s flexible kernel-based approach handles both cases. We also compared against MMD [21] and the Frechet distance [22, 23], with NPLM proving competitive or superior in most settings.
Bonus: Re-Discovering the Higgs Boson in Real LHC Data
As a fun bonus demonstration, we applied AutoSciDACT to real proton-proton collision data from the CMS experiment at the LHC, searching for evidence of the Higgs boson in the four-lepton decay channel – the same channel that contributed to the Higgs discovery in 2012 [24, 25].
Using only a contrastive encoder trained on simulated background processes (with no knowledge of the Higgs) and a basic pre-selection targeting events with four leptons in the final state, we ran NPLM on CMS Open Data and examined which events received the highest anomaly scores. The figure below tells the story: the horizontal axis shows the four-lepton invariant mass \(m_{4\ell}\), the vertical axis shows the NPLM anomaly score, and the most anomalous events in real data (red points) cluster near 120–130 GeV – exactly where the Higgs boson lives at 125 GeV. The horizontal dashed lines show example thresholds on the anomaly score; applying a tight cut would isolate a handful of Higgs-like events from the background, effectively “re-discovering” the Higgs without being told to look for it.
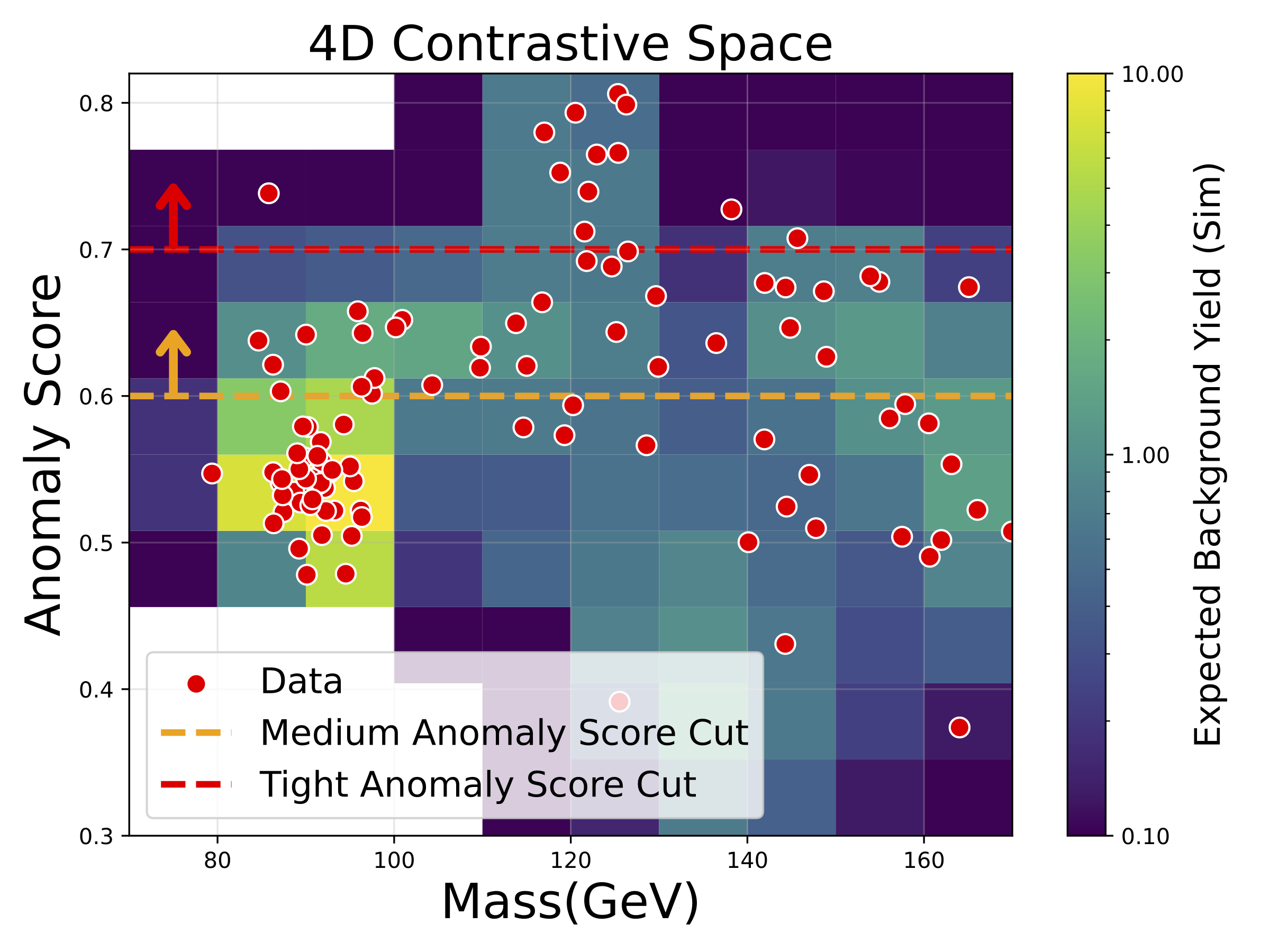
An important caveat: this analysis uses a very small dataset from a single Higgs decay channel, and we do not account for systematic uncertainties from domain shift between simulation and real data. The resulting p-values do not meet the bar for a statistically significant finding, let alone a discovery – even the original CMS analysis [24] achieved only ~2-sigma expected significance in this channel alone, claiming discovery only by combining multiple channels. We present this result purely as a demonstration of AutoSciDACT’s ability to home in on real anomalous structure in collider data, not as a standalone measurement.
Why This Matters: Bridging AI and the Scientific Method
AutoSciDACT isn’t just another anomaly detector. It’s an attempt to formalize and automate a core part of the scientific method itself: the cycle of observation, hypothesis formation, and rigorous statistical testing. By decoupling domain knowledge (which enters only through labels and architecture choices in the pre-training phase) from the analysis (which is entirely signal-agnostic), it provides a framework that can be dropped into essentially any scientific domain where:
- You have labeled examples of known phenomena (from simulations, expert annotation, or prior measurements).
- You want to search for unknown deviations in new data.
- You need a statistically rigorous claim about what you find.
This describes a huge swath of modern science – from particle physics to astronomy to biology to materials science. The key insight is that the statistical machinery for hypothesis testing, refined over decades in particle physics, can be made universal when paired with the right representation learning.
Limitations and Future Directions
We’re transparent about what AutoSciDACT can’t yet do. The pipeline currently assumes the reference distribution faithfully represents the true background, which may not hold when there are domain shifts between simulation and real data. It relies on label quality for pre-training, and aggressive compression to 4 dimensions necessarily sacrifices some expressivity. Future work will address these gaps – incorporating systematic uncertainties [7], handling domain shift, and exploring higher-dimensional embeddings where warranted.
This work is supported by the National Science Foundation under Cooperative Agreement PHY-2019786 (The NSF AI Institute for Artificial Intelligence and Fundamental Interactions, IAIFI). The full paper is available at OpenReview.
References
[1] P. Khosla et al. “Supervised Contrastive Learning.” NeurIPS 33, 18661–18673, 2020.
[2] T. Chen et al. “A Simple Framework for Contrastive Learning of Visual Representations.” ICML, 1597–1607, 2020.
[3] R. T. D’Agnolo, A. Wulzer. “Learning New Physics from a Machine.” Phys. Rev. D 99(1), 015014, 2019.
[4] R. T. D’Agnolo et al. “Learning Multivariate New Physics.” Eur. Phys. J. C 81(1), 89, 2021.
[5] J. Neyman, E. S. Pearson. “On the Problem of the Most Efficient Tests of Statistical Hypotheses.” Phil. Trans. R. Soc. A 231, 289–337, 1933.
[6] M. Letizia et al. “Learning New Physics Efficiently with Nonparametric Methods.” Eur. Phys. J. C 82(10), 879, 2022.
[7] G. Grosso et al. “Goodness of Fit by Neyman-Pearson Testing.” SciPost Physics 16(5), 123, 2024.
[8] G. Grosso, M. Letizia. “Multiple Testing for Signal-Agnostic Searches for New Physics with Machine Learning.” Eur. Phys. J. C 85(1), 4, 2025.
[9] H. Qu et al. “Particle Transformer for Jet Tagging.” arXiv:2202.03772, 2022.
[10] A. Vaswani et al. “Attention Is All You Need.” NeurIPS 30, 2017.
[11] J. Aasi et al. (LIGO Scientific Collaboration). “Advanced LIGO.” Class. Quantum Grav. 32, 074001, 2015.
[12] K. He et al. “Deep Residual Learning for Image Recognition.” CVPR, 770–778, 2016.
[13] E. Marx et al. “Machine-Learning Pipeline for Real-Time Detection of Gravitational Waves from Compact Binary Coalescences.” Phys. Rev. D 111(4), 042010, 2025.
[14] I. Zingman et al. “NAFLD Pathology and Healthy Tissue Samples.” 2022. Available at https://osf.io/gqutd/.
[15] M. Tan, Q. V. Le. “EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks.” ICML, 6105–6114, 2019.
[16] I. Zingman et al. “Learning Image Representations for Anomaly Detection: Application to Discovery of Histological Alterations in Drug Development.” Medical Image Analysis 92, 103067, 2024.
[17] S. Stevens et al. “BioCLIP: A Vision Foundation Model for the Tree of Life.” arXiv:2311.18803, 2023.
[18] K. Lee et al. “A Simple Unified Framework for Detecting Out-of-Distribution Samples and Adversarial Attacks.” NeurIPS 31, 2018.
[19] G. Kasieczka et al. “The LHC Olympics 2020: A Community Challenge for Anomaly Detection in High Energy Physics.” Rep. Prog. Phys. 84(12), 124201, 2021.
[20] R. Raikman et al. “A Neural Network-Based Search for Unmodeled Transients in LIGO-Virgo-KAGRA’s Third Observing Run.” arXiv:2412.19883, 2024.
[21] A. Gretton et al. “A Kernel Two-Sample Test.” JMLR 13, 723–773, 2012.
[22] D. C. Dowson, B. Landau. “The Frechet distance between multivariate normal distributions.” J. Multivariate Anal. 12(3), 450–455, 1982.
[23] M. Heusel et al. “GANs trained by a two time-scale update rule converge to a local Nash equilibrium.” NeurIPS 30, 2017.
[24] S. Chatrchyan et al. (CMS Collaboration). “Observation of a New Boson at a Mass of 125 GeV with the CMS Experiment at the LHC.” Phys. Lett. B 716, 30–61, 2012.
[25] G. Aad et al. (ATLAS Collaboration). “Observation of a New Particle in the Search for the Standard Model Higgs Boson with the ATLAS Detector at the LHC.” Phys. Lett. B 716, 1–29, 2012.